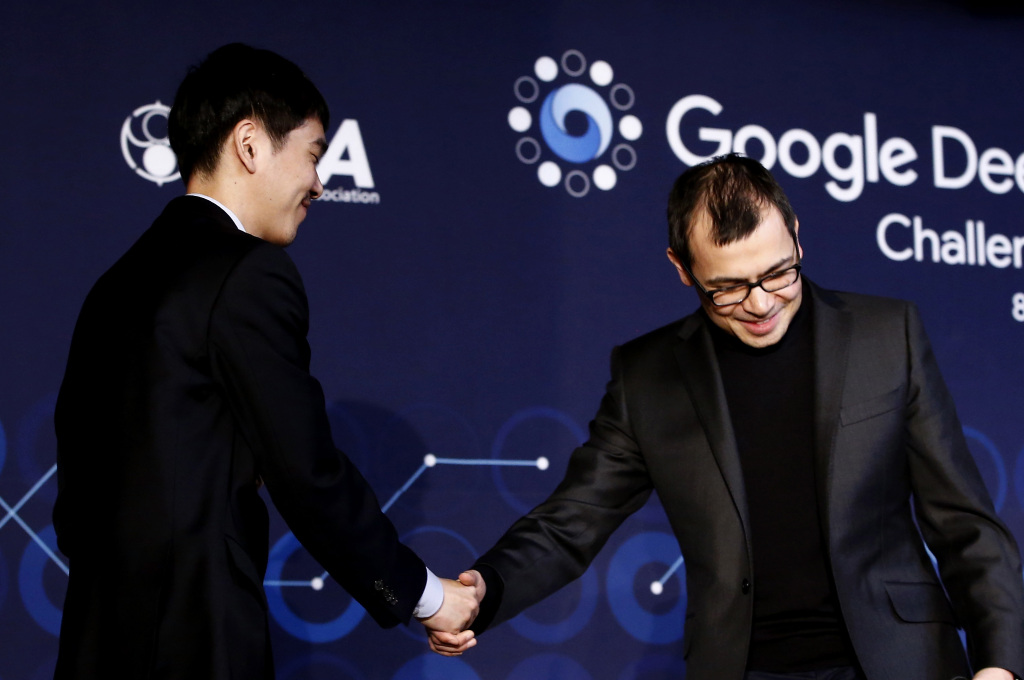
It is March 13, 2016. Two men, dressed in winter coats and woolen hats to defend against the frigid night air, walk side-by-side through the crowded streets of downtown Seoul. Locked in animated conversation, they seem oblivious to the pulsating neon enticements of the surrounding dumpling houses and barbeque joints. They are visitors, come to South Korea on a mission, the culmination of years of effort—and they have just succeeded.
This is a celebratory stroll. What they have achieved will cement their places in the annals of computer science: They have built a piece of artificial intelligence software able to play the ancient strategy game Go so expertly that it handily defeated the world’s top player, Lee Sedol. Now the two men are discussing their next goal, their conversation captured by a documentary film crew shadowing them.
“I’m telling you, we can solve protein folding,” Demis Hassabis, says to his walking companion, David Silver. “That’s like, I mean, it’s just huge. I am sure we can do that now. I thought we could do that before, but now we definitely can do it.” Hassabis is the co-founder and chief executive officer of DeepMind, the London-based A.I. company that built AlphaGo. Silver is the DeepMind computer scientist who led the AlphaGo team.
Four years later, DeepMind has just accomplished what Hassabis broached in that nocturnal amble: It has created an A.I. system that can predict the complex shapes of proteins down to an atom’s-width accuracy from the genetic sequences that encode them. With this achievement, DeepMind has completed an almost 50-year-old scientific quest. In 1972, in his Nobel Prize acceptance speech, chemist Christian Anfinsen postulated that DNA alone should fully determine the final structure a protein takes. It was a remarkable conjecture. At the time, not a single genome had been sequenced yet. But Anfinsen’s theory launched an entire sub-field of computational biology with the goal of using complex mathematics, instead of empirical experiments, to model proteins.
DeepMind’s achievement with Go was important—but it had little concrete impact outside the relatively cliquish worlds of Go and computer science. Solving protein folding is different: it could prove transformative to much of humanity. Proteins are the basic building blocks of life and the mechanism behind most biological processes. Being able to predict their structure could revolutionize our understanding of disease and lead to new, more targeted pharmaceuticals for everything from cancer to Alzheimer’s disease. It will likely accelerate the time it takes to bring new medicines to market, potentially shaving years and hundreds of millions of dollars in cost from drug development, and potentially saving lives as a result.

The new method pioneered by DeepMind is already yielding results in the fight against SARS-CoV-2, the virus that causes COVID-19. What follows is the story of how a company best known for playing games came to unlock one of biology’s greatest secrets.
Building blocks with elusive shapes
“Proteins are the main machines of the cell,” Ian Holmes, a professor of bioengineering at the University of California at Berkeley, says. “And the structure and shape of them is crucial to how they operate.” Small “pockets” within the lattice of molecules that make up the protein are where various chemical reactions take place. If you can find a chemical that will bind to one of these pockets, then that substance can be used as a drug—to either disable or accelerate a biological process. Bioengineers can also create entirely new proteins never before seen in nature with unique therapeutic properties. “If we could tap into the power of proteins and rationally engineer them to any purpose, then we could build these remarkable self-assembling machines that could do things for us,” Holmes says.
But to be sure the protein will do what you want, it’s important to know its shape.
Proteins consist of chains of amino acids, often compared to beads on a string. The recipe for which beads to string in what order is encoded in DNA. But the complex physical shape the completed chain will take is extremely difficult to predict from those simple genetic instructions. Amino acid chains collapse—or fold—into a structure based on electrochemical rules of attraction and repulsion between molecules. The resulting shapes frequently resemble abstract sculptures formed from tangles of cord and ribbon: pleated banderoles joined to Mobius strip-like curlicues and looping helixes. In the 1960s, Cyrus Levinthal, a physicist and molecular biologist, determined that there were so many plausible shapes a protein might assume that it would take longer than the known age of the universe to arrive at the correct structure by randomly trying combinations—and yet, the protein folds itself in milliseconds. This observation has become known as Levinthal’s Paradox.
Until now, the only way to know a protein’s structure with near certainty was through a method known as X-ray crystallography. As the name implies, this involves turning solutions of millions of proteins into crystals, a chemical process that is itself tricky. X-rays are then fired at these crystals, allowing a scientist to work backwards from the diffraction patterns they make to build up a picture of the protein itself. Oh, and not just any X-rays: For many proteins, the X-rays need to be produced by a massive, stadium-sized circular particle accelerator called a synchrotron.
The process is expensive and time-consuming: it takes about 12 months and approximately $120,000 to determine a single protein’s structure with X-ray crystallography, according to one estimate from researchers at the University of Toronto. There are over 200 million known proteins, with about 30 million more being discovered every year, and yet fewer than 200,000 of these have had their structures mapped with X-ray crystallography or other experimental methods. “Our level of ignorance is growing rapidly,” says John Jumper, a computational physicist who is now a senior researcher at DeepMind and leads its protein folding team.
Over the past fifty years, ever since Christian Anfinsen’s famous speech, scientists have tried speed up the analysis of protein structure by using complex mathematical models run on high-powered computers. “What you do is essentially try to create a digital twin of the protein in your computer, and then try to manipulate it,” says John Moult, a professor of cell biology and molecular genetics at the University of Maryland and a pioneer in using mathematical algorithms to predict protein structures from their DNA sequences. The problem is, these predicted folding patterns were frequently wrong, failing to match the structures scientists found through X-ray crystallography. In fact, until about ten years ago, few models were able to accurately predict more than about a third of a large protein’s shape.
Some protein folding simulations also take up tremendous amounts of computing power. In the year 2000, researchers created a “citizens science” project called Fold@home in which people could donate the idle processing capacity of their personal computers and game consoles to run a protein folding simulation. All those devices, chained together through the Internet, created one of the world’s most powerful virtual supercomputers. The hope was that this would allow researchers to escape Levinthal’s Paradox—to speed up the time it would take to hit upon the accurate protein structures through random trial-and-error. The project, which is still running, has provided data for more than 225 scientific papers on proteins implicated in a number of diseases.
But despite having access to so much processing power, Fold@home is still mired in Levinthal’s Paradox: it is trying to find a protein structure by searching through all possible permutations. The holy grail of protein folding is to skip this laborious search and to instead discover elusive patterns that link a proteins’ DNA sequence to its structure—allowing a computer to take a radical shortcut, leaping directly from genetics to the correct shape.
Games with a serious purpose
Demis Hassabis’s interest in protein folding began, as many of Hassabis’s passions do, with a game. Hassabis is a former chess prodigy, a master by the time he was 13 and at one time ranked second in the world for his age. His love of chess fed a fascination with two things: game design and the inner mechanisms of his own mind. He began working for a video games company while still in high school and, after studying computer science at the University of Cambridge, founded his own computer games startup, Elixir Studios, in 1998.
Despite producing two award-winning games, Elixir eventually sold off its intellectual property and shut down, and Hassabis went on to get a Ph.D. in cognitive neuroscience from University College London. By then, he had already embarked on the crusade that would lead him to co-found DeepMind in 2010: the creation of artificial general intelligence—software capable of learning to perform many disparate tasks as well or better than people. DeepMind’s lofty goal, Hassabis once said, was “to solve intelligence, and then use it to solve everything else.” Hassabis already had an inkling that protein folding just might be one of those first “everything elses.”
Hassabis was doing a post-doc at the Massachusetts Institute of Technology in 2009 when he heard about an online game called Foldit. Foldit was designed by researchers at the University of Washington and, like Fold@home, it was a “citizens science” project for protein folding. But instead of yoking together idle microchips, Foldit was designed to harness idle brains.
Foldit is a puzzle-like game in which human players, without any knowledge of biology, compete to fold proteins, earning points for creating shapes that are plausible. Researchers then analyze the highest-scoring designs to see if they can help complete unsolved protein structures. The game has attracted tens of thousands of players and, in a number of documented cases, produced better protein structures than protein-folding computer algorithms. “I thought it was fascinating from the standpoint of, can we use the addictiveness of games and the joy of them, and in the background not only are they having fun, but they are doing something useful for science,” Hassabis says.
But there was another reason Foldit would continue to capture Hassabis’s imagination. Games are a particularly good arena for a kind of A.I. training called reinforcement learning. This is where software learns from experience, essentially by trial and error, to get better at a task. In a computer game, software can experiment endlessly, playing over and over again, improving gradually until it reaches superhuman skill, without causing any real-world harm. Games also have ready-made and unambiguous ways to tell if a particular action or set of actions is effective: points and wins. Those metrics provide a very clear way to benchmark performance—something that doesn’t exist for many real-world problems, where the most effective move may be far more ambiguous and the entire concept of “winning” may not apply.
DeepMind was founded largely on the promise of combining reinforcement learning with a kind of A.I. called deep learning. Deep learning is A.I. based on neural networks—a kind of software loosely based on how the human brain works. In this case, instead of networks of actual nerve cells, the software has a bunch of virtual neurons, arranged in a hierarchy where an initial input layer takes in some data, applies a weighting to it, and passes it along to the middle layers, which do the same in turn, until it is eventually passed to an output layer that sums up all the weighted signals and uses that to produce a result. The network adjusts these weights until it can produce a desired outcome—such as accurately identifying photos of cats or winning a game of chess. It’s called “deep learning” not because the insights it produces are necessarily profound—although they can be—but because the network consists of many layers and so can be said to have depth.
DeepMind’s initial success came in using this “deep reinforcement learning” to create software that taught itself to play classic Atari computer games, such as Pong, Breakout and Space Invaders, at superhuman levels. It was this achievement that helped get DeepMind noticed by big technology firms, including Google, which bought it for a reported £400 million (more than $600 million at the time) in 2014. It then turned its attention to Go, eventually creating the system AlphaGo, which defeated Sedol in 2016. DeepMind went on to create a more general version of that system, called AlphaZero, that could learn to play almost any two-player, turn-based game in which players have perfect information (so there is no element of chance or hidden information, such as face down cards or hidden positions) at superhuman levels. Last year, it also built a system that could beat top human professional e-sports players at the highly complex real-time strategy game Starcraft 2.

But Hassabis says he always saw the company’s work with games as a way to perfect A.I. methods so they could be applied to real world challenges—especially in science. “Games are just a training ground, but a training ground for what exactly? For creating new knowledge,” he says.
DeepMind is not a traditional business, with products and customers. Instead, it is essentially a research lab that tries to advance the frontiers of artificial intelligence. Many of the methods it develops, it publishes openly for anyone to use or build upon. But some of its advances are useful for its sister company, Google.
DeepMind has a whole team of engineers and scientists that help Google incorporate cutting-edge A.I. into its products. DeepMind’s technology has found its way into everything from Google Maps to the company’s digital assistant to the system that helps manage battery power on Android phones. Google pays DeepMind for this help, and Alphabet, its parent company, continues to absorb the additional losses that DeepMind generates. Those are not insignificant: The company lost £470 million in 2018 (about $510 million at the time), the last year for which its annual financial statements are publicly available through the U.K. business registry Companies House.
But DeepMind, which now employs more than 1,000 people, also has a whole other division that works only on scientific applications of A.I. It is headed by Pushmeet Kohli, a 39-year-old native of India, who worked on A.I. research for Microsoft before joining DeepMind. He says that DeepMind’s aim is to try to solve “root node” problems—data science-speak for saying it wants to take on issues that are fundamental to unlocking many different scientific avenues. Protein folding is one of these root nodes, Kohli says.
‘The Olympics of protein-folding’
In 1994, at a time when many scientists were first starting to use sophisticated computer algorithms to try to predict how proteins would fold, Moult, the University of Maryland biologist, decided to create a competition that could provide an unbiased way of assessing which of these algorithms was best. He called this competition the Critical Assessment of Protein Structure Prediction (CASP, for short), and it has been held biennially ever since.
It works like this: The Protein Structure Prediction Center, the organization that runs CASP and which is funded through the U.S. National Institute of General Medical Sciences, persuades researchers who do X-ray crystallography and other empirical studies to provide it with protein structures that have not yet been published anywhere, asking them to refrain from making the structures public until after the CASP competition. CASP then gives the DNA sequences of these proteins to the contestants, who use their algorithms to predict the protein’s structure. CASP then judges how close the predictions are to the actual structure the X-ray crystallographers and experimentalists found. The algorithms are then ranked by their average performance across all the proteins. “I call it the Olympics of protein folding,” Hassabis says. And, in 2016, shortly after AlphaGo beat Sedol, DeepMind set out to win the gold medal.
DeepMind established a small, crack team of a half-dozen machine learning researchers and engineers to work on the problem. “It’s part of our philosophy that we start with generalists,” Hassabis says. The company does not suffer from a lack of brain power. “Ex-physicists, ex-biologists, we just have them lying around generally,” Hassabis says with a wry smile. “They never know when their previous expertise suddenly is going to become useful.” Eventually the team grew to about 20 people.
Still, DeepMind decided it would be helpful to have at least one true protein-folding expert onboard. It found one in John Jumper. Skinny, with a mop of asymmetrically styled brown hair, Jumper is a boyish 35, and looks a bit like the bass guitarist in a late-1990s high school garage band. He earned a master’s degree in theoretical condensed matter physics from Cambridge before going on to work at D.E. Shaw Research in New York City, an independent research lab founded by hedge fund billionaire David Shaw. The lab specializes in computational biology, including the simulation of proteins. Jumper later got his PhD. in computational biophysics from the University of Chicago, studying under Karl Freed and Tobin Sosnick, two scientists known for advances in protein fold modeling. “I had heard this rumor that DeepMind was interested in protein problems,” he says. He applied and got the job.
Hassabis’s and the DeepMind team’s first instinct was that protein folding could be solved in exactly the same way as Go—with deep reinforcement learning. But this proved problematic: For one thing, there were even more possible fold configurations than there are moves in Go. More importantly, DeepMind had mastered Go in large part by getting its A.I. system, AlphaGo, to play games against itself. “There isn’t quite the right analogy for that because protein folding is not a two-player game,” Hassabis says. “You’re sort of playing against Nature.”

DeepMind soon established that there was a simpler way of making progress using a kind of A.I. training known as supervised deep learning. This is the sort of A.I. used in most business applications: From an established set of data inputs and corresponding outputs, a neural network learns how to match a given input to a given output. In this case, DeepMind had the protein structures—currently about 170,000 of them—that are publicly available in the Protein Data Bank (PDB), a public repository of all known three-dimensional protein shapes and their genetic sequences, to use as training data.
Some biologists had already used supervised deep learning to predict how proteins would fold. But the best of these A.I. systems were only right about 50% of the time, which wasn’t particularly helpful to biologists or medical researchers—especially since, for a protein whose structure was unknown, they had no way of determining whether a particular prediction was correct.
One promising technique rested on the idea that proteins can be grouped into families based on their evolutionary history. And within these families, it is possible to find pairs of amino acids that are distant from one another in a DNA sequence, yet seem to mutate at the same time. This phenomenon, which is called “co-evolution,” is helpful because co-evolved proteins are likely to be in contact within the protein’s folded structure. Jinbo Xu, a scientist at the Toyota Technological Institute in Chicago, pioneered using deep learning on this co-evolutionary data to predict amino acid contacts. The approach is a bit like finding just the dots in a connect-the-dots game. Scientists still had to use other software to try to figure out the lines between those dots—and often they got this wrong. Sometimes they didn’t even get the dots rights.
For the 2018 CASP competition, DeepMind took these basic ideas about co-evolution and contact prediction but added two important twists. First, rather than trying to determine if two amino acids were in contact, a binary output (either the pair is in contact or isn’t), it decided to ask the algorithm to predict the distance between all the amino acid pairs in the protein.
To most molecular biologists, such an approach seemed counterintuitive—although Xu, to his credit, had independently proposed a similar method. After all, it was contact that mattered most. But to DeepMind’s deep learning experts it was immediately obvious that distance was a much better metric for a neural network to work on, Kohli says. “It is just a fundamental part of deep learning that if you have some uncertainty associated with a decision, it is much better to have the neural network incorporate that uncertainty and decide what to do about it,” he says. Distance, unlike contact, was a richer piece of information the network could adjust and play with.
The other twist DeepMind came up with was a second neural network that predicted the angles between amino acid pairs. With these two factors—distance and angles—DeepMind’s algorithm was able to work out a rough outlines of a protein’s likely structure. It then used a different, non-A.I. algorithm to refine this structure. Putting these components together into a system it called AlphaFold, DeepMind crushed the competition in the 2018 CASP (called CASP13 because it was the 13th of the biennial contests). On the hardest set of 43 proteins in the competition, AlphaFold got the highest score on 25 of them. The next closest team scored highest on just three. The results shook the entire field: if there had been any doubt about whether deep learning methods were the most promising way to crack the protein-folding problem, AlphaFold ended them.
Going back to the whiteboard
Still, DeepMind was nowhere close to Hassabis’s goal: solving the protein-folding problem. AlphaFold was fairly inaccurate almost half the time. And, of the 104 protein targets in CASP13, it achieved results that were as good as X-ray crystallography in only about three cases. “We didn’t just want to be the best at this according to CASP, we wanted to be good at this. We actually want a system that matters to biologists,” Jumper says.
No sooner had the CASP 2018 results been announced than DeepMind redoubled its efforts: Jumper was put in charge of expanded team. Rather than simply trying to build on AlphaFold, making incremental improvements, the team went back to the whiteboard and started to brainstorm radically different ideas that they hoped would be able to bring the software closer to the kind of accuracy X-ray crystallography yielded.
What followed, Jumper says, was one of scariest and most depressing periods of the entire project: nothing worked. “We spent three months not getting any better than our CASP13 results and starting to really panic,” he says. But then, a few of the things the researchers were trying produced a slight improvement—and within six months the system was notably better than the original AlphaFold. This pattern would continue throughout the next two years, Jumper says: three months of nothing, followed by three months of rapid progress, followed by yet another plateau.
Hassabis says a similar pattern had occurred with previous DeepMind projects, including its work on Go and the complex, real-time strategy video game Starcraft 2. The company’s management strategy for overcoming this, he says, is to alternate between two different ways of working. The first, which Hassabis calls “strike mode,” involves pushing the team as hard as possible to wring every ounce of performance out of an existing system. Then, when the gains from the all-out effort seem to be exhausted, he shifts gears into what he calls “creative mode.” During this period, Hassabis no longer presses the team on performance—in fact, he tolerates and even expects some temporary declines—in order to give the researchers and engineers the space to tinker with new ideas and try novel approaches. “You want to encourage as many crazy ideas as possible, brainstorming,” he says. This often leads to another leap forward in performance, allowing the team to switch back into strike mode.
A big birthday present
On Nov. 21 of 2019, Kathryn Tunyasuvunakool, a researcher at DeepMind who works on the protein folding team, turned 30. The day would prove to be memorable for another reason too. Tunyasuvunakool, who has a Ph.D. in computational biology from the University of Oxford, was the person on the team in charge of developing new test sets for the protein folding A.I., now dubbed AlphaFold 2, that DeepMind was developing for the 2020 CASP competition. That morning, when she turned on her office computer, she received an assessment of the system’s predictions on a batch of about 50 protein sequences—all of them only recently added to the Protein Data Bank. She did a double take. AlphaFold 2 had been improving, but on this set of proteins the results were startlingly good—predicting the structure in many cases to within 1.5 angstroms, a distance equivalent to a tenth of a nanometer, or about the width of an atom.

Tunyasuvunakool, who calls herself “the team’s pessimist,” says her first response was not elation, but nausea. “I was feeling quite scared,” she says. The results were so good she was certain she had made a mistake—that when she was preparing the test set, she must have inadvertently allowed several proteins that the A.I. had already seen in the training data to slip in. That would have allowed AlphaFold 2 to essentially cheat, easily predicting the exact structure. Tunyasuvunakool recalls sitting in DeepMind’s cafeteria overlooking London’s St. Pancras Station and drinking cup after cup of herbal tea in an effort to calm herself. She and other team members then spent the rest of that day and late into the evening, and several days more, sitting at their workstations, painstakingly combing through AlphaFold 2’s training data to try to find the mistake.
There wasn’t one. In fact, the new system had made a giant leap forward in performance. AlphaFold 2 was completely different from its predecessor. Rather than an assemblage of components—one to predict the distance between amino acids and another to forecast the angles, with a third piece of software to tie them together —the A.I. now used a single neural network to reason directly from the DNA sequence. While the system still took in evolutionary information—figuring out if the protein in question had a likely common ancestor to others it had seen before, and scrutinizing the alignment between the target protein’s DNA sequence and other known sequences—it no longer needed explicit data about which amino acid pairs evolved together. “Instead of providing more information, we actually provided less,” Jumper says. The system was free to draw its own insights about when ancestry might determine a portion of the protein’s shape and when it might depart more radically from that heritage. In other words, it developed a kind of intuition based on its experience, in much the same way a veteran human scientist might.
At the heart of the new system was a mechanism called “attention.” Attention, as the name implies, is a way to get a deep learning system to focus on a certain set of inputs and weigh those more heavily. For a cat identification system, for instance, the system might learn to pay attention to the shape of the ears and also learn to look for evidence of whiskers near the nose. Jumper compares what AlphaFold 2 does to the process of solving a jigsaw puzzle where “you can snap together certain pieces and be pretty sure of it, and then what you end up with are different local islands of solution, and then you figure out how to join these up.” The middle of the network, Jumper says, has learned to reason about geometry and space and how to join up those amino acid pairs it thinks are close together based on its analysis of the DNA sequences.
DeepMind trained AlphaFold 2 on 128 “tensor processing cores,” the number-crunching brains found on 16 special computer chips engineered for deep learning that Google designed and uses in its data centers, running continuously for what the company says was a few weeks. (These 128 specialized A.I. cores are about equivalent to 100 to 200 of the powerful graphics processing chips that deliver eye-popping animation on an Xbox or Playstation.) Once trained, the system can take a DNA sequence and spit out a complete structure prediction “in a matter of days,” the company says.
Among AlphaFold 2’s advantages over its predecessor is a confidence gauge: The system produces a score for how sure it is of its own predictions for each amino acid in a structure. This metric is crucial if AlphaFold 2 is going to be useful to biologists and medical researchers who will need to know when they can reasonably rely on the model and when to have more caution.
Despite the stunning test results, DeepMind was still not certain how good AlphaFold 2 was. But they got an important clue when the coronavirus pandemic struck. In March of this year, AlphaFold 2 was able to predict the structure for six understudied proteins associated with SARS-CoV-2, the virus that causes COVID-19, one of which scientists have since confirmed using an empirical method called cryogenic electron microscopy. It was a powerful glimpse of the kind of real-world impact DeepMind hopes AlphaFold 2 will soon have. (To read more about AlphaFold 2’s role in the fight against COVID-19, click here.)
An astonishing result
The CASP competition takes place between May and August. The Protein Structure Prediction Center releases batches of target proteins and contestants then submit their structure predictions for evaluation. The rankings for this year’s competition were announced on Nov. 30.
Each prediction is scored using a metric called “global distance test total score,” or GDT for short, that in effect looks at how close, in angstroms, it is to a structure obtained by empirical methods such as X-ray crystallography or electron microscope. A score of 100 is perfect, but anything at 90 or above is considered equivalent to the empirical methods, Moult, the CASP director, says. The proteins are also classed into groups based on how difficult the CASP organizers think it is to get the structure.
When Moult saw AlphaFold 2’s results he was incredulous. Like Tunyasuvunakool months earlier, his initial thought was that there might be a mistake. Maybe some of the protein sequences in the competition had been published before? Or maybe DeepMind had somehow managed to get hold of a cache of unpublished data?
As a test, he asked Andrei Lupas, director of the department of protein evolution at the Max Planck Institute for Developmental Biology in Tuebingen, Germany, to conduct an experiment. Lupas would ask AlphaFold 2 to predict a structure that he knew for certain had never been seen before because Lupas had never been able to work out from X-ray crystallography what a key piece of the protein looked like. For almost a decade, Lupas had puzzled over this missing link, but the correct shape had eluded him. Now, with AlphaFold’s prediction as a guide, Lupas says, he went back to the X-ray data. “The correct structure just fell out within half an hour,” he says. “It was astonishing.”
Since DeepMind’s success in 2018’s CASP, many academic researchers have flocked to deep learning techniques. As a result, the rest of the field’s performance has improved: On a median difficulty target, the other competitors now have an average best prediction GDT of 75, up 10 points from two years ago. But there was no comparison to AlphaFold 2: It scored a median 92 GDT across all proteins, and even on the most difficult proteins it achieved a median score of 87 GDT. Moult says AlphaFold 2’s predictions are “on par with empirical methods,” such as X-ray crystallography. That conclusion lead CASP to make a momentous declaration on Monday, Nov. 30: The 50-year-old protein-folding problem had been solved.
Venki Ramakrishnan, a Nobel prize-winning structural biologist who is also the current president of The Royal Society, Britain’s most prestigious scientific body, says AlphaFold2 “represents a stunning advance” in protein-folding. With AlphaFold 2, expensive and time consuming empirical analysis with methods like X-ray crystallography and electron microscopes may become a thing of the past.
Janet Thornton, an expert in protein structure and former director of the European Molecular Biology Laboratory’s European Bioinformatics Institute, says that DeepMind’s breakthrough will allow scientists to map the entire human “proteome”—all the proteins found within the human body. Currently only a quarter of these proteins have been used as targets for drugs, but having the structure for the rest would create vast opportunities for the development of new therapies. She also says the A.I. software could enable protein engineering that might aid in sustainability efforts, allowing scientists to potentially create new crop strains that provide more nutritional value per acre of land planted, and also possibly allowing for the advent of enzymes that could digest plastic.
For now though, the question remains about how exactly DeepMind will make AlphaFold 2 available. Hassabis says the company is committed to ensuring the software can “make the maximal positive societal impact.” But he says it has not yet determined how to do that, saying only that it will make an announcement sometime next year. Hassabis also tells Fortune that DeepMind in considering how it might be able to build a commercial product or partnership around the system. “This should be hugely useful for the drug discovery process and therefore Big Pharma,” he says. But exactly what form this commercial offering will take, he says, has not yet been decided either.
A commercial venture would be marked departure for DeepMind, which, since its sale to Alphabet, has not had to worry about generating revenue. The company briefly set up a division called DeepMind Health that was working with the U.K.’s National Health Service on an app that could identify hospital patients who were at risk of developing acute kidney injury. But the effort became embroiled in a controversy after news reports revealed DeepMind’s hospital partner had violated the U.K. data protection laws by giving the company access to millions of patients’ medical records. In 2019, DeepMind Health was formally absorbed into a new Google health division. At the time, DeepMind said cleaving off its health effort would allow it to remain true to its research roots without the distraction of having to build a commercial unit that might replicate areas, such as data security and customer support, where Google already had expertise.
Of course, if DeepMind launch a commercial product, it would not be the first A.I. research company to do so: OpenAI, the San Francisco-based research company that is perhaps DeepMind’s closest rival, has become increasingly business-oriented. Last year, OpenAI launched its first commercial product, an interface that lets companies use an A.I. that composes long passages of coherent text from a short, human-written prompt. The business value of that A.I., called GPT-3, remains unproven, while DeepMind’s AlphaFold 2 could have an immediate bottom line impact for a pharmaceutical company or biotechnology startup. At a time when antitrust regulators are probing Alphabet, having a viable commercial product could be a good insurance policy for DeepMind in the event it ever loses the unconditional support of its deep-pocketed parent in some future break-up of the Googleplex.
One thing is certain: DeepMind isn’t done with protein folding. The CASP competition was set up around predicting the structure of single proteins. But in biology and medicine, it is usually protein interactions that researchers really care about. How does one protein bind with another or with a particular small molecule? Exactly how does an enzyme break a protein apart? The problem of predicting these interactions and bindings will likely become the primary focus of future CASP competitions, Moult says. And Jumper says DeepMind plans to work on those challenges next.
Reverberations from AlphaFold 2’s success are certain to be felt in areas far removed from protein folding, too, encouraging others to apply deep learning to big scientific questions: finding new sub-atomic particles, probing the secrets of dark matter, mastering nuclear fusion or creating room-temperature superconductors. DeepMind has an active effort already underway on astrophysics, Kohli says. Facebook’s A.I. researchers just launched a deep learning project aimed at finding new chemical catalysts. Protein-folding is the first foundational scientific mystery to fall to the power of artificial intelligence. It won’t be the last.
More health care and Big Pharma coverage from Fortune:
- Why it’s hard to process 250,000 COVID deaths
- Your employees are not okay: How to handle mental health at work during a pandemic
- You’re not crazy—you really hunger for social contact, scientists say
- The Fortune/IBM Watson Health 50 Top Cardiovascular Hospitals
- Lockdown, superspreader, unprecedented: 2020 has changed the English language, for good




